Circular RNAs (circRNAs) are a class of RNA molecules characterized by a covalent circular structure formed through a non-canonical 3′ to 5′ end-joining process known as back-splicing (BS). They can regulate gene expression through diverse mechanisms, including acting as transcriptional regulators, sponging miRNAs, modulating mRNA stability, and scaffolding proteins. Therefore, identifying their target RNA interactions is a prerequisite for understanding their full functional repertoire.
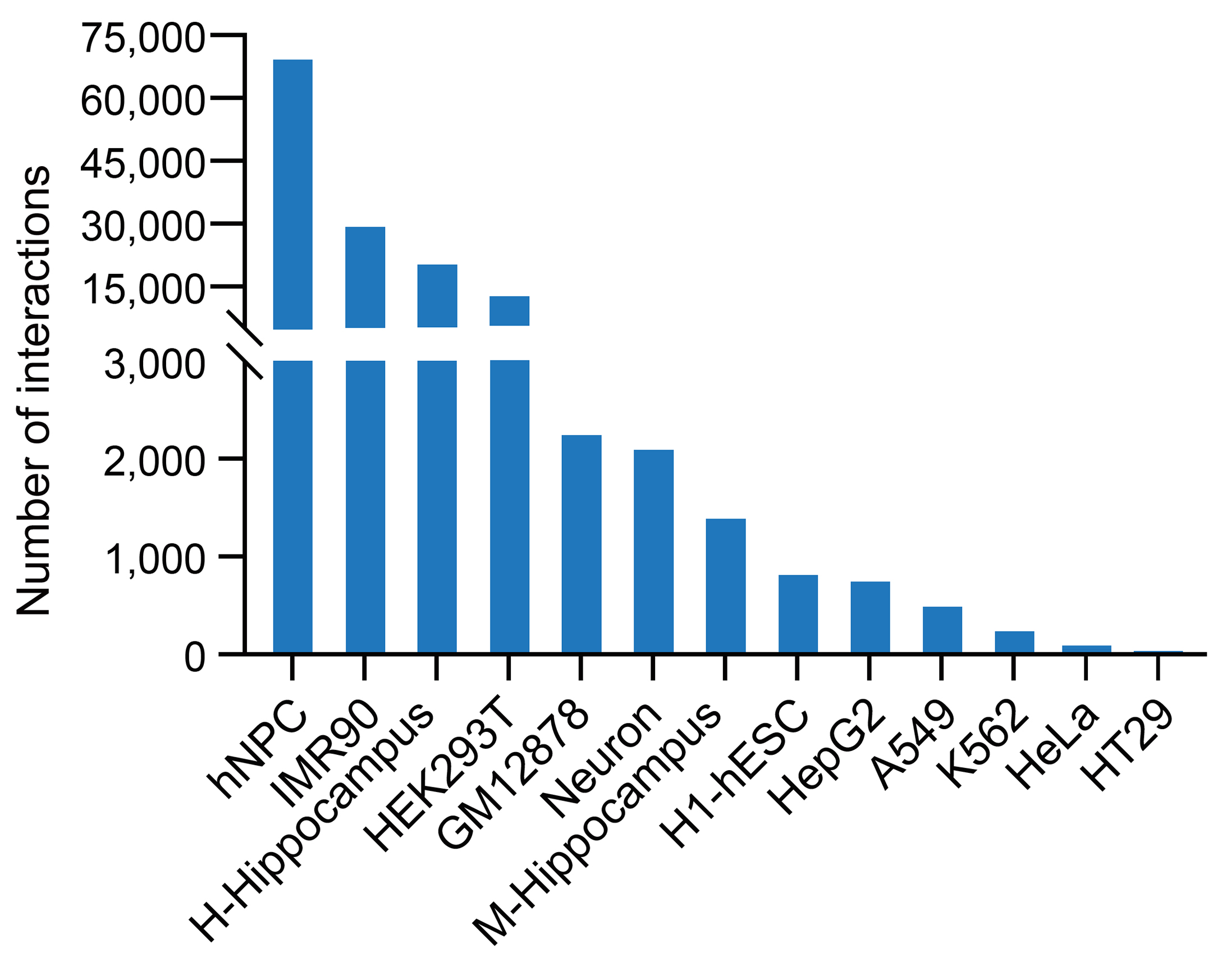
In the CircTarget database, we systematically identified circRNA-target RNA interactions using datasets from RIC-seq, KARR-seq, LIGR-seq, PARIS, and SPLASH across 13 cell lines and hippocampal tissues of human and mouse. Given the low abundance of circRNAs and their sequence overlap with linear RNAs, we first identified interactions using chimeric reads spanning their unique back-splicing junctions (BSJs), yielding 9,935 high-confidence pairs. For circRNAs with extreme circularization ratios (circular-to-linear ratio ≥ 0.85, e.g., CDR1as), we expanded our analysis to include all chimeric reads mapped to their full-length transcripts, identifying an additional 122,582 interactions.
Each entry in CircTarget is comprehensively annotated, including circRNA basic information, disease associations, interaction details, predicted hybrid duplex structures, and disease-linked variants.
chimeric reads (n = 9,935)
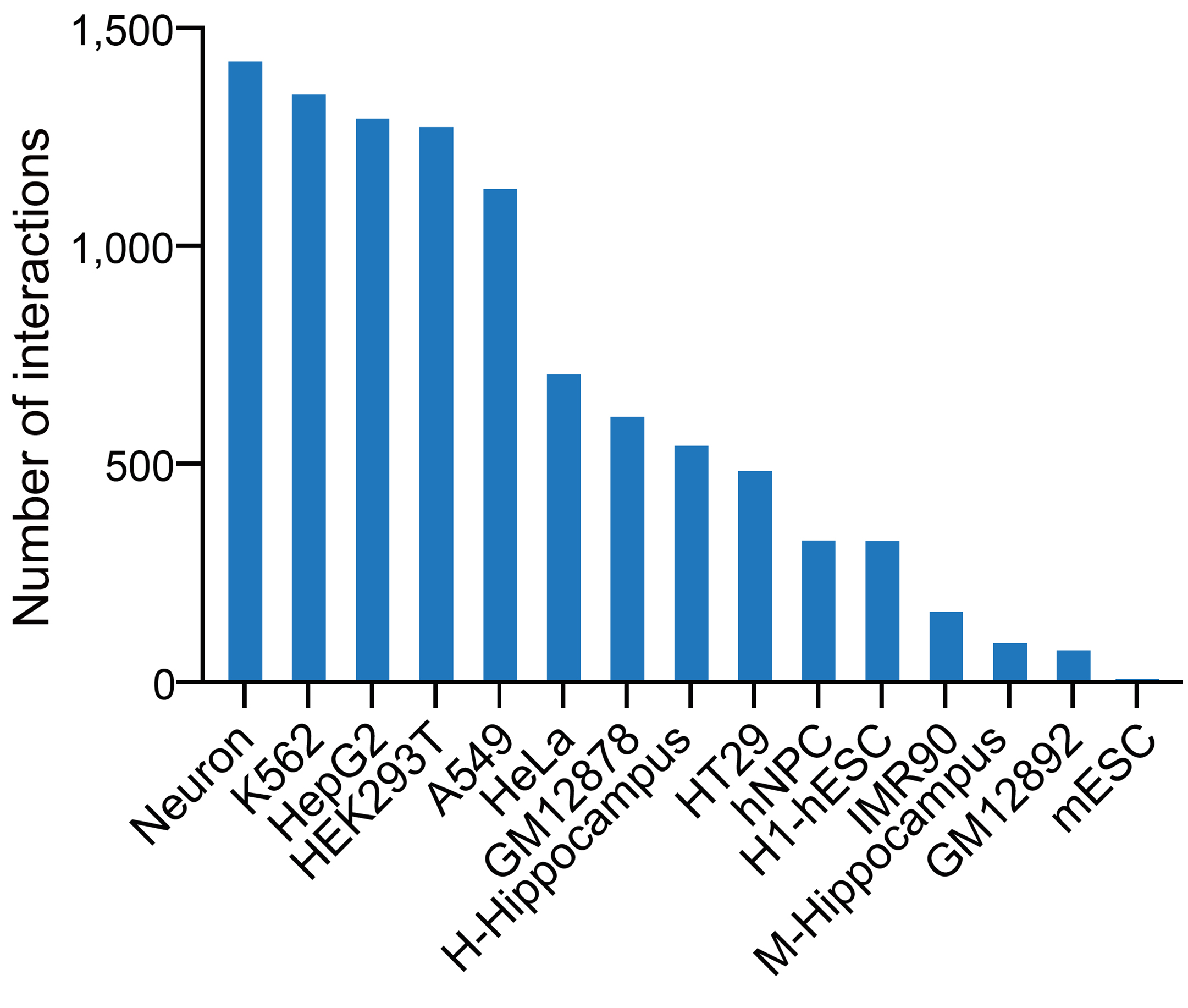
chimeric reads (n = 122,582)